This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Parkinson's Disease and PINK1
|
Parkinson's Disease (PD) is the second most common neurodegenerative disease. PD affects 1-2% of the population; about 1 million people in the United States. [1] Parkinson's Disease is a movement disorder caused by the degeneration of dopaminergic neurons in the substantia nigra. The average age of onset is 60 years old. If an individual is diagnosed between ages 20-40, it is considered early-onset Parkinson's Disease. [1]
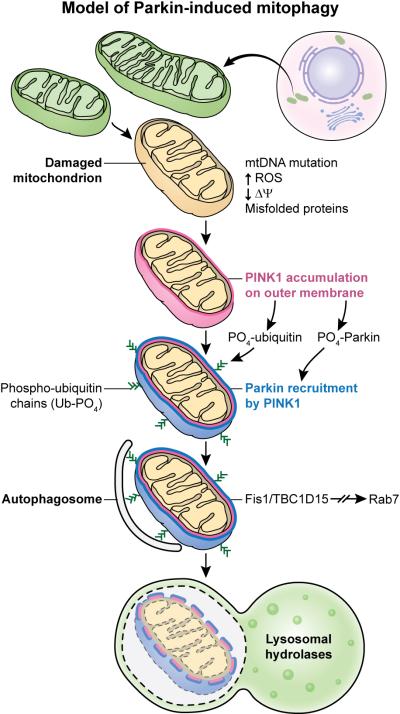
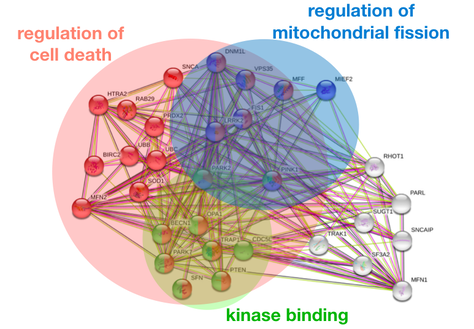
The exact etiology of Parkinson's disease in unknown, but there are several genes that have been identified as risk factors , including PINK1. PINK1 is an autosomal recessive gene associated with early-onset Parkinson's disease that is involved in the mitophagy pathway. PINK1 recruits and phosphorylates Parkin, which recruits other factors to phagocytize the damaged mitochondrion (1). PINK1 localizes in the mitochondria, which makes sense, as its primary molecular function and biological processes are phosphorylating Parkin in the mitophagy pathway. The PINK1 protein interaction network further demonstrates PINK1's role in the mitophagy pathway, as the majority of the protein interactions are with other proteins involved mitochondrial regulation and regulation of cell death. Because mitophagy of damaged mitochondria is an important process in almost all living species, PINK1 is well-conserved with the serine/threonine protein kinase domain present in organisms from Humans to C. elegans to Arabidopsis. |
Gap in Knowledge
Goals and Hypothesis
My primary goal is to better understand how regulation of PINK1 kinase activity contributes to mitochondrial regulation and neurodegeneration. My long-term goal of this research is to understand how regulation of PINK1 contributes to regulation of mitophagy so that effective treatments can be made for Parkinson’s disease patients. I will test the hypothesis that PINK1 kinase activity can be regulated by altering autophosphorylation sites in PINK1.
Model Organism
|
I will use zebrafish as a model organism because the region of the zebrafish brain that contains dopaminergic neurons is strikingly similar to the human ascending midbrain DA neurons, and development of this region is believed to be very similar to human development (2). Mitochondrial dysfunction can be viewed in zebrafish as an increase in mitochondrial size in cells.
Throughout my study, I will be comparing mutant PINK1 zebrafish to wildtype zebrafish. |
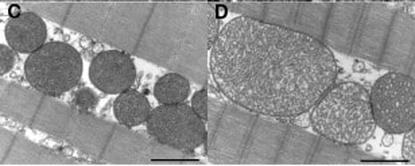
Figure C. shows healthy mitochondria. Figure D. shows enlarged, dysfunctional mitochondria.
|
Aim 1
Aim 1: Identify conserved autophosphorylation sites of PINK1 crucial for normal mitophagy pathway.
Approach: First, I will use BLAST to find homologs of PINK1. I will then align protein sequences using Clustal Omega to identify conserved autophosphorylation sites that have been found in insect PINK1 among the homologs. I will then mutate the well conserved autophosphorylation sites with CRISPR-Cas9 using zebrafish as a model to understand how these mutations affect the mitophagy pathway. I will then screen the mutated zebrafish for the increased mitochondrial size phenotype, as well as dopaminergic death at different stages of life.
Approach: First, I will use BLAST to find homologs of PINK1. I will then align protein sequences using Clustal Omega to identify conserved autophosphorylation sites that have been found in insect PINK1 among the homologs. I will then mutate the well conserved autophosphorylation sites with CRISPR-Cas9 using zebrafish as a model to understand how these mutations affect the mitophagy pathway. I will then screen the mutated zebrafish for the increased mitochondrial size phenotype, as well as dopaminergic death at different stages of life.
Rationale: Autophosphorylation sites have been identified in insect PINK1 that are believed to autophosphorylate before PINK1 can phosphorylate Parkin. Screening of zebrafish with the induced gene mutations from those conserved sites should result in zebrafish with dysfunctional mitochondria that are increased in size, which will help explain what mutations in PINK1 do to the mitophagy pathway.
Hypothesis: I hypothesize that specific autophosphorylation sites in PINK1 correlate with normal mitophagy pathway and decreased dopaminergic death.
Hypothesis: I hypothesize that specific autophosphorylation sites in PINK1 correlate with normal mitophagy pathway and decreased dopaminergic death.
Aim 2
Aim 2: Identify small molecules that rescue PINK1 mutant phenotypes.
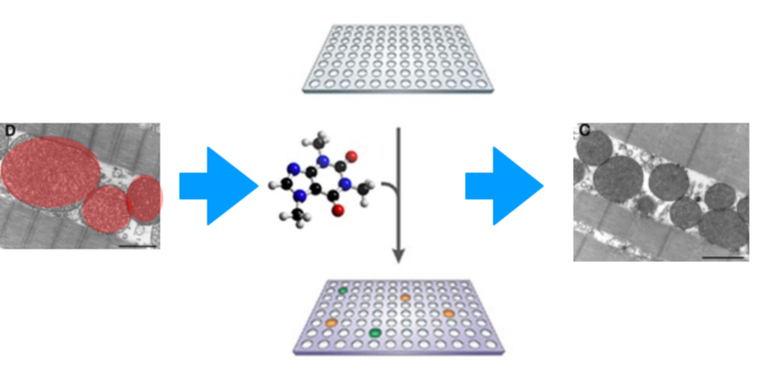
Approach: I will perform a high-throughput chemical genomic screen on wild type and PINK1 mutant zebrafish from Aim 1 at different ages. This will be done using a diversity-oriented library to identify small molecules that rescue the PINK1 mutant dysfunctional mitochondria and dopaminergic death.
Approach: I will perform a high-throughput chemical genomic screen on wild type and PINK1 mutant zebrafish from Aim 1 at different ages. This will be done using a diversity-oriented library to identify small molecules that rescue the PINK1 mutant dysfunctional mitochondria and dopaminergic death.
Rationale: Treating mutant PINK1 zebrafish with the identified small molecules will restore the mitophagy pathway and restore wild type function of PINK1.
Hypothesis: I hypothesize that small molecules that can rescue the PINK1 mutant phenotypes will restore the normal mitophagy pathway and decrease dopaminergic death.
Hypothesis: I hypothesize that small molecules that can rescue the PINK1 mutant phenotypes will restore the normal mitophagy pathway and decrease dopaminergic death.
Aim 3
Aim 3: Determine differences in protein interaction complexes required for normal mitophagy pathway between mutant and wildtype zebrafish.
Approach: I will use autophosphorylation site mutants from Aim 1 with dysfunctional mitochondria. For mutant and wildtype groups, PINK1 fragment-containing complexes will be copurified with the affinity-tagged PINK1-fragment proteins and analyzed using mass spectrometry. I will then compare the mutant and wildtype zebrafish to each other and create protein-protein interaction webs. I will find the gene ontology of interaction proteins using Panther.
Approach: I will use autophosphorylation site mutants from Aim 1 with dysfunctional mitochondria. For mutant and wildtype groups, PINK1 fragment-containing complexes will be copurified with the affinity-tagged PINK1-fragment proteins and analyzed using mass spectrometry. I will then compare the mutant and wildtype zebrafish to each other and create protein-protein interaction webs. I will find the gene ontology of interaction proteins using Panther.
Rationale: In normal individuals, PINK1 works with other proteins in the mitophagy pathway. I want to elucidate if mutation of the autophosphorylation sites affects the protein-protein interaction network responsible for mitophagy.
Hypothesis: I hypothesize that mutated PINK1 will have reduced protein interactions with mitochondrial regulation and apoptosis regulation proteins.
Hypothesis: I hypothesize that mutated PINK1 will have reduced protein interactions with mitochondrial regulation and apoptosis regulation proteins.
Future Directions
|
Early-onset Parkinson's disease is extremely debilitating and currently has no cure. In the future, I would like to look more into differences in protein interactions between mutant and wildtype PINK1 and try to understand why the differences exist. I also think it would be useful to find some sort of target for reporters of active PINK1, after it has autophosphorylated.I would also like to mutate Parkin and see how those mutations affect its interaction with PINK1.
|
References:
1. de Rijk MC, Breteler MM, Graveland GA, Ott A, Grobbee DE, van der Meche FG, Hofman A. Prevalence of Parkinson's disease in the elderly: the Rotterdam Study. Neurology. 1995;45:2143–2146. https://www.ncbi.nlm.nih.gov/pubmed/8848182
2. Xi, Y., Noble, S., & Ekker, M. (2011). Modeling neurodegeneration in zebrafish. Current neurology and neuroscience reports, 11(3), 274–282. doi:10.1007/s11910-011-0182-2
1. de Rijk MC, Breteler MM, Graveland GA, Ott A, Grobbee DE, van der Meche FG, Hofman A. Prevalence of Parkinson's disease in the elderly: the Rotterdam Study. Neurology. 1995;45:2143–2146. https://www.ncbi.nlm.nih.gov/pubmed/8848182
2. Xi, Y., Noble, S., & Ekker, M. (2011). Modeling neurodegeneration in zebrafish. Current neurology and neuroscience reports, 11(3), 274–282. doi:10.1007/s11910-011-0182-2